!pip install mlcroissant -qSTAC to GeoCroissant Conversion
This notebook converts metadata from STAC Items and Collections into GeoCroissant JSON-LD format.

GeoCroissant is a geospatial extension of MLCommons’ Croissant metadata standard for ML-ready dataset descriptions.
STAC to GeoCroissant Mapping
| STAC Field | GeoCroissant Field |
|---|---|
id |
@id, name |
type |
Detected (Item vs Collection) |
description |
description |
license |
license |
bbox (Item) |
spatialCoverage |
extent.spatial (Coll.) |
spatialCoverage |
properties.datetime |
temporalCoverage, datePublished |
extent.temporal (Coll.) |
temporalCoverage, datePublished |
properties.proj:epsg |
geocr:coordinateReferenceSystem |
properties.proj:wkt2 |
geocr:coordinateReferenceSystem |
assets[].gsd |
geocr:spatialResolution |
assets[].bands |
geocr:bandConfiguration |
links |
url, references |
assets |
distribution |
providers |
creator |
Import Required Libraries
Convert STAC to GeoCroissant
This converter handles both STAC Items (type: “Feature”) and STAC Collections.
import json
from datetime import datetime
import re
def sanitize_name(name):
return re.sub(r"[^a-zA-Z0-9_\-]", "-", name)
def ensure_semver(version):
if not version:
return "1.0.0"
if version.startswith("v"):
version = version[1:]
parts = version.split(".")
if len(parts) == 2:
parts.append("0")
return ".".join(parts[:3])
def extract_crs_from_wkt2(wkt2_string):
"""Extract EPSG code or return WKT2 description."""
if not wkt2_string:
return None
# Check if this is a known projected coordinate system by looking for PROJCS
if "PROJCS" in wkt2_string or "PROJECTION" in wkt2_string:
# This is a projected CRS - try to identify it
if "Albers" in wkt2_string:
return "Custom Albers Equal Area projection"
elif "UTM" in wkt2_string:
# Try to extract UTM zone
zone_match = re.search(r'UTM[^\d]*(\d+)', wkt2_string, re.IGNORECASE)
if zone_match:
return f"UTM Zone {zone_match.group(1)}"
return "UTM projection"
else:
return "Custom projection (see proj:wkt2)"
# For geographic coordinate systems, try to find EPSG code at the DATUM or GEOGCS level
# But be careful not to pick up unit codes like EPSG:9122
epsg_match = re.search(r'AUTHORITY\["EPSG","(\d+)"\]', wkt2_string)
if epsg_match:
epsg_code = epsg_match.group(1)
# Skip common unit codes
if epsg_code not in ["9122", "9001"]:
return f"EPSG:{epsg_code}"
return None
def stac_to_geocroissant(stac_dict):
# Detect if this is a STAC Item (Feature) or Collection
stac_type = stac_dict.get("type", "").lower()
is_item = stac_type == "feature"
is_collection = stac_type == "collection"
if is_item:
print("Detected STAC Item (Feature)")
elif is_collection:
print("Detected STAC Collection")
else:
print(f"Unknown STAC type: {stac_type}, treating as Collection")
dataset_id = stac_dict.get("id")
# For Items, use id directly; for Collections, prefer title
if is_item:
name = sanitize_name(dataset_id or "UnnamedItem")
else:
name = sanitize_name(stac_dict.get("title", dataset_id or "UnnamedDataset"))
version = ensure_semver(stac_dict.get("version", "1.0.0"))
croissant = {
"@context": {
"@language": "en",
"@vocab": "https://schema.org/",
"citeAs": "cr:citeAs",
"column": "cr:column",
"conformsTo": "dct:conformsTo",
"cr": "http://mlcommons.org/croissant/",
"geocr": "http://mlcommons.org/croissant/geo/",
"rai": "http://mlcommons.org/croissant/RAI/",
"dct": "http://purl.org/dc/terms/",
"sc": "https://schema.org/",
"data": {"@id": "cr:data", "@type": "@json"},
"examples": {"@id": "cr:examples", "@type": "@json"},
"dataBiases": "cr:dataBiases",
"dataCollection": "cr:dataCollection",
"dataType": {"@id": "cr:dataType", "@type": "@vocab"},
"extract": "cr:extract",
"field": "cr:field",
"fileProperty": "cr:fileProperty",
"fileObject": "cr:fileObject",
"fileSet": "cr:fileSet",
"format": "cr:format",
"includes": "cr:includes",
"isLiveDataset": "cr:isLiveDataset",
"jsonPath": "cr:jsonPath",
"key": "cr:key",
"md5": "cr:md5",
"parentField": "cr:parentField",
"path": "cr:path",
"personalSensitiveInformation": "cr:personalSensitiveInformation",
"recordSet": "cr:recordSet",
"references": "cr:references",
"regex": "cr:regex",
"repeated": "cr:repeated",
"replace": "cr:replace",
"samplingRate": "cr:samplingRate",
"separator": "cr:separator",
"source": "cr:source",
"subField": "cr:subField",
"transform": "cr:transform"
},
"@type": "sc:Dataset",
"@id": dataset_id,
"name": name,
"description": stac_dict.get("description", ""),
"version": version,
"license": stac_dict.get("license", "CC-BY-4.0"),
"conformsTo": [
"http://mlcommons.org/croissant/1.0",
"http://mlcommons.org/croissant/geo/1.0"
]
}
# Add citeAs (recommended property)
if is_collection:
collection_name = stac_dict.get("title", name)
croissant["citeAs"] = f"@misc{{{dataset_id}, title={{{collection_name}}}, year={{{datetime.utcnow().year}}}}}"
else:
# For items, try to get collection info
collection_id = stac_dict.get("collection", dataset_id)
croissant["citeAs"] = f"@misc{{{collection_id}, title={{{name}}}, year={{{datetime.utcnow().year}}}}}"
if "sci:citation" in stac_dict:
croissant["citeAs"] = stac_dict["sci:citation"]
croissant["citation"] = stac_dict["sci:citation"]
if stac_dict.get("providers"):
provider = stac_dict["providers"][0]
croissant["creator"] = {
"@type": "Organization",
"name": provider.get("name", "Unknown"),
"url": provider.get("url", "")
}
# Handle 'self' URL
for link in stac_dict.get("links", []):
if link.get("rel") == "self":
croissant["url"] = link.get("href")
break
# Handle other STAC references
references = []
for link in stac_dict.get("links", []):
rel = link.get("rel")
href = link.get("href")
if not href or rel == "self":
continue
name_map = {
"root": "STAC root catalog",
"parent": "STAC parent catalog",
"items": "STAC item list",
"about": "GitHub Repository",
"predecessor-version": "Previous version",
"http://www.opengis.net/def/rel/ogc/1.0/queryables": "Queryables"
}
references.append({
"@type": "CreativeWork",
"url": href,
"name": name_map.get(rel, rel),
"encodingFormat": link.get("type", "application/json")
})
if references:
croissant["references"] = references
# === Spatial Coverage ===
# For STAC Items: use bbox directly
# For Collections: use extent.spatial.bbox
bbox = None
if is_item and "bbox" in stac_dict:
bbox = stac_dict["bbox"]
elif not is_item:
spatial = stac_dict.get("extent", {}).get("spatial", {}).get("bbox")
if spatial and spatial[0]:
bbox = spatial[0]
if bbox and len(bbox) >= 4:
# STAC bbox format: [west, south, east, north]
# GeoShape box format: "south west north east"
croissant["spatialCoverage"] = {
"@type": "Place",
"geo": {
"@type": "GeoShape",
"box": f"{bbox[1]} {bbox[0]} {bbox[3]} {bbox[2]}"
}
}
# === Temporal Coverage ===
# For STAC Items: use properties.start_datetime/end_datetime or properties.datetime
# For Collections: use extent.temporal.interval
if is_item:
props = stac_dict.get("properties", {})
start_dt = props.get("start_datetime")
end_dt = props.get("end_datetime")
datetime_val = props.get("datetime")
if start_dt and end_dt:
croissant["temporalCoverage"] = f"{start_dt}/{end_dt}"
croissant["datePublished"] = start_dt.split("T")[0]
elif datetime_val:
croissant["temporalCoverage"] = datetime_val
croissant["datePublished"] = datetime_val.split("T")[0]
# Use created_datetime if available
if "created_datetime" in props and "datePublished" not in croissant:
croissant["datePublished"] = props["created_datetime"].split("T")[0]
else:
# Collection extent
temporal = stac_dict.get("extent", {}).get("temporal", {}).get("interval")
if temporal and temporal[0]:
start, end = temporal[0][0], temporal[0][1]
if start and end:
croissant["temporalCoverage"] = f"{start}/{end}"
elif start:
croissant["temporalCoverage"] = start
if start:
croissant["datePublished"] = start.split("T")[0]
if "datePublished" not in croissant:
croissant["datePublished"] = datetime.utcnow().isoformat().split("T")[0]
# Asset-level distribution
croissant["distribution"] = []
all_bands = [] # Collect band info from all assets
for key, asset in stac_dict.get("assets", {}).items():
file_object = {
"@type": "cr:FileObject",
"@id": key,
"name": key,
"description": asset.get("description", asset.get("title", "")),
"contentUrl": asset.get("href"),
"encodingFormat": asset.get("type", "application/octet-stream")
}
# Only add checksums if they exist
if "checksum:multihash" in asset:
file_object["sha256"] = asset["checksum:multihash"]
elif "file:checksum" in asset:
file_object["sha256"] = asset["file:checksum"]
if "checksum:md5" in asset:
file_object["md5"] = asset["checksum:md5"]
# Croissant requires at least one checksum (md5 or sha256)
# If STAC doesn't provide checksums, add a placeholder
if "sha256" not in file_object and "md5" not in file_object:
file_object["sha256"] = "PLACEHOLDER-CHECKSUM-REQUIRED"
print(f" Warning: No checksum found for asset '{key}'. Added placeholder.")
print(f" Please compute SHA256 for: {asset.get('href', 'N/A')}")
# Extract band information if present
if "bands" in asset:
bands = asset["bands"]
if bands and len(bands) > 0:
band_config = {
"@type": "geocr:BandConfiguration",
"geocr:totalBands": len(bands),
"geocr:bandNamesList": []
}
# Try to extract band names from band metadata
for i, band in enumerate(bands):
if "name" in band:
band_config["geocr:bandNamesList"].append(band["name"])
else:
band_config["geocr:bandNamesList"].append(f"Band{i+1}")
file_object["geocr:bandConfiguration"] = band_config
all_bands.extend(bands)
# Extract spatial resolution from gsd if present
if "gsd" in asset and "geocr:spatialResolution" not in croissant:
croissant["geocr:spatialResolution"] = {
"@type": "QuantitativeValue",
"value": asset["gsd"],
"unitText": "m"
}
croissant["distribution"].append(file_object)
# item_assets as fileSet templates
if "item_assets" in stac_dict:
croissant["fileSet"] = []
for key, asset in stac_dict["item_assets"].items():
file_obj = {
"@type": "cr:FileObject",
"@id": key,
"name": key,
"description": asset.get("description", asset.get("title", "")),
"encodingFormat": asset.get("type", "application/octet-stream")
}
file_set = {
"@type": "cr:FileSet",
"name": f"Template for {key}",
"includes": [file_obj]
}
croissant["fileSet"].append(file_set)
# Keywords
if "keywords" in stac_dict:
croissant["keywords"] = stac_dict["keywords"]
# === CRS and Spatial Resolution ===
# For Items: check properties for proj:epsg or proj:wkt2
# For Collections: check summaries
if is_item:
props = stac_dict.get("properties", {})
# Extract CRS
if "proj:epsg" in props:
croissant["geocr:coordinateReferenceSystem"] = f"EPSG:{props['proj:epsg']}"
elif "proj:wkt2" in props:
crs_result = extract_crs_from_wkt2(props["proj:wkt2"])
if crs_result:
croissant["geocr:coordinateReferenceSystem"] = crs_result
# Spatial resolution already extracted from asset gsd above
else:
# Collection summaries
summaries = stac_dict.get("summaries", {})
if "proj:epsg" in summaries:
epsg_codes = summaries["proj:epsg"]
if epsg_codes and len(epsg_codes) > 0:
croissant["geocr:coordinateReferenceSystem"] = f"EPSG:{epsg_codes[0]}"
# Spatial resolution from summaries
if "gsd" in summaries and "geocr:spatialResolution" not in croissant:
gsd_values = summaries["gsd"]
if gsd_values and len(gsd_values) > 0:
croissant["geocr:spatialResolution"] = {
"@type": "QuantitativeValue",
"value": gsd_values[0],
"unitText": "m"
}
# Note: renders, summaries, stac_extensions, and stac_version are STAC-specific
# and not part of the GeoCroissant vocabulary, so they are not included
if "deprecated" in stac_dict:
croissant["isLiveDataset"] = not stac_dict["deprecated"]
# Report unmapped fields
mapped_keys = {
"id", "type", "links", "title", "assets", "extent",
"license", "version", "providers", "description", "sci:citation",
"renders", "summaries", "stac_extensions", "stac_version", "deprecated",
"item_assets", "keywords", "bbox", "geometry", "properties", "collection"
}
extra_fields = {k: v for k, v in stac_dict.items() if k not in mapped_keys}
print("\n\033[1mUnmapped STAC Fields:\033[0m")
if extra_fields:
for k, v in extra_fields.items():
print(f"- {k}: {type(v).__name__}")
else:
print("None")
return croissant
# === Main Runner ===
if __name__ == "__main__":
# Load STAC Collection JSON
with open("stac.json") as f:
stac_data = json.load(f)
# Convert to GeoCroissant
croissant_json = stac_to_geocroissant(stac_data)
# Save GeoCroissant JSON-LD
with open("croissant.json", "w") as f:
json.dump(croissant_json, f, indent=2)
print("\nGeoCroissant conversion complete. Output saved to 'croissant.json'")Detected STAC Item (Feature)
Warning: No checksum found for asset 'cog'. Added placeholder.
Please compute SHA256 for: s3://nasa-maap-data-store/file-staging/nasa-map/icesat2-boreal-v2.1/agb/0038382/boreal_agb_2020_202411261732637661_0038382.tif
Warning: No checksum found for asset 'training_data_csv'. Added placeholder.
Please compute SHA256 for: s3://nasa-maap-data-store/file-staging/nasa-map/icesat2-boreal-v2.1/agb/0038382/boreal_agb_2020_202411261732637661_0038382_train_data.csv
Unmapped STAC Fields:
None
GeoCroissant conversion complete. Output saved to 'croissant.json'Preview Generated GeoCroissant JSON
Inspect the converted metadata structure and values.
import json
# Read and print the metadata
with open('croissant.json', 'r') as f:
metadata = json.load(f)
# Pretty print the metadata
print(json.dumps(metadata, indent=2)){
"@context": {
"@language": "en",
"@vocab": "https://schema.org/",
"citeAs": "cr:citeAs",
"column": "cr:column",
"conformsTo": "dct:conformsTo",
"cr": "http://mlcommons.org/croissant/",
"geocr": "http://mlcommons.org/croissant/geo/",
"rai": "http://mlcommons.org/croissant/RAI/",
"dct": "http://purl.org/dc/terms/",
"sc": "https://schema.org/",
"data": {
"@id": "cr:data",
"@type": "@json"
},
"examples": {
"@id": "cr:examples",
"@type": "@json"
},
"dataBiases": "cr:dataBiases",
"dataCollection": "cr:dataCollection",
"dataType": {
"@id": "cr:dataType",
"@type": "@vocab"
},
"extract": "cr:extract",
"field": "cr:field",
"fileProperty": "cr:fileProperty",
"fileObject": "cr:fileObject",
"fileSet": "cr:fileSet",
"format": "cr:format",
"includes": "cr:includes",
"isLiveDataset": "cr:isLiveDataset",
"jsonPath": "cr:jsonPath",
"key": "cr:key",
"md5": "cr:md5",
"parentField": "cr:parentField",
"path": "cr:path",
"personalSensitiveInformation": "cr:personalSensitiveInformation",
"recordSet": "cr:recordSet",
"references": "cr:references",
"regex": "cr:regex",
"repeated": "cr:repeated",
"replace": "cr:replace",
"samplingRate": "cr:samplingRate",
"separator": "cr:separator",
"source": "cr:source",
"subField": "cr:subField",
"transform": "cr:transform"
},
"@type": "sc:Dataset",
"@id": "boreal_agb_2020_202411261732637661_0038382",
"name": "boreal_agb_2020_202411261732637661_0038382",
"description": "",
"version": "1.0.0",
"license": "CC-BY-4.0",
"conformsTo": [
"http://mlcommons.org/croissant/1.0",
"http://mlcommons.org/croissant/geo/1.0"
],
"citeAs": "@misc{icesat2-boreal-v2.1-agb, title={boreal_agb_2020_202411261732637661_0038382}, year={2026}}",
"url": "https://stac.maap-project.org/collections/icesat2-boreal-v2.1-agb/items/boreal_agb_2020_202411261732637661_0038382",
"references": [
{
"@type": "CreativeWork",
"url": "https://stac.maap-project.org/collections/icesat2-boreal-v2.1-agb",
"name": "collection",
"encodingFormat": "application/json"
},
{
"@type": "CreativeWork",
"url": "https://stac.maap-project.org/collections/icesat2-boreal-v2.1-agb",
"name": "STAC parent catalog",
"encodingFormat": "application/json"
},
{
"@type": "CreativeWork",
"url": "https://stac.maap-project.org/",
"name": "STAC root catalog",
"encodingFormat": "application/json"
}
],
"spatialCoverage": {
"@type": "Place",
"geo": {
"@type": "GeoShape",
"box": "72.7215138641646 111.821583637516 73.8622051330815 115.626213868067"
}
},
"temporalCoverage": "2020-01-01T00:00:00Z/2020-12-31T23:59:59Z",
"datePublished": "2020-01-01",
"distribution": [
{
"@type": "cr:FileObject",
"@id": "cog",
"name": "cog",
"description": "Gridded predictions of aboveground biomass (Mg/ha)",
"contentUrl": "s3://nasa-maap-data-store/file-staging/nasa-map/icesat2-boreal-v2.1/agb/0038382/boreal_agb_2020_202411261732637661_0038382.tif",
"encodingFormat": "image/tiff; application=geotiff; profile=cloud-optimized",
"sha256": "PLACEHOLDER-CHECKSUM-REQUIRED",
"geocr:bandConfiguration": {
"@type": "geocr:BandConfiguration",
"geocr:totalBands": 2,
"geocr:bandNamesList": [
"Band1",
"Band2"
]
}
},
{
"@type": "cr:FileObject",
"@id": "training_data_csv",
"name": "training_data_csv",
"description": "Tabular training data with latitude, longitude, and biomass observations",
"contentUrl": "s3://nasa-maap-data-store/file-staging/nasa-map/icesat2-boreal-v2.1/agb/0038382/boreal_agb_2020_202411261732637661_0038382_train_data.csv",
"encodingFormat": "text/csv",
"sha256": "PLACEHOLDER-CHECKSUM-REQUIRED"
}
],
"geocr:spatialResolution": {
"@type": "QuantitativeValue",
"value": 30,
"unitText": "m"
},
"geocr:coordinateReferenceSystem": "Custom Albers Equal Area projection"
}Validate with mlcroissant
Validate the generated GeoCroissant JSON-LD against the specification using the mlcroissant CLI.
!mlcroissant validate --jsonld=croissant.jsonimport json
# Load your croissant.json
with open("croissant.json", "r") as f:
croissant = json.load(f)
# Print all reference links
print("\nReferenced STAC Links:")
for ref in croissant.get("references", []):
name = ref.get("name")
url = ref.get("url")
fmt = ref.get("encodingFormat")
print(f" - {name}: {url} ({fmt})")
Referenced STAC Links:
- collection: https://stac.maap-project.org/collections/icesat2-boreal-v2.1-agb (application/json)
- STAC parent catalog: https://stac.maap-project.org/collections/icesat2-boreal-v2.1-agb (application/json)
- STAC root catalog: https://stac.maap-project.org/ (application/json)import json
import rasterio
from rasterio.plot import show
import matplotlib.pyplot as plt
import numpy as np
# Load croissant.json to get raster URL
with open("croissant.json", "r") as f:
croissant_data = json.load(f)
# Extract the raster URL from distribution
raster_url = None
for dist in croissant_data.get("distribution", []):
content_url = dist.get("contentUrl", "")
if content_url.endswith(".tif") or "tif" in content_url:
raster_url = content_url
raster_name = dist.get("name", "unnamed")
print(f"Found raster: {raster_name}")
break
if not raster_url:
print("No raster file found in croissant.json")
else:
# Prepare URL for rasterio based on protocol
if raster_url.startswith("s3://"):
cog_url = raster_url.replace("s3://", "/vsis3/")
elif raster_url.startswith("http://") or raster_url.startswith("https://"):
cog_url = f"/vsicurl/{raster_url}"
else:
cog_url = raster_url
print(f"Opening: {cog_url}")
with rasterio.Env(AWS_NO_SIGN_REQUEST="YES"):
with rasterio.open(cog_url) as src:
print(f"CRS: {src.crs}")
print(f"Bounds: {src.bounds}")
print(f"Shape: {src.height} x {src.width}")
print(f"Bands: {src.count}")
# Read band statistics
for i in range(1, src.count + 1):
band = src.read(i, masked=True)
mask = band.mask
data = band.data
valid_pixels = np.count_nonzero(~mask)
print(f"\nBand {i} Stats:")
if valid_pixels > 0:
print(f" Min: {data[~mask].min():.2f}")
print(f" Max: {data[~mask].max():.2f}")
print(f" Mean: {data[~mask].mean():.2f}")
else:
print(" No valid data")
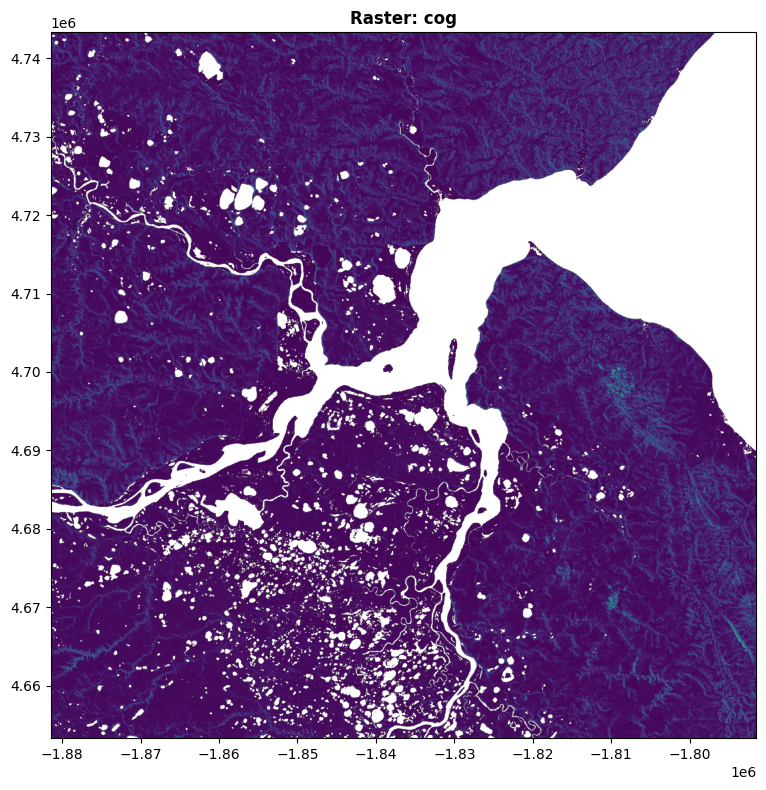
# Display the raster
fig, ax = plt.subplots(figsize=(12, 8))
show(src, ax=ax, title=f"Raster: {raster_name}")
plt.tight_layout()
plt.show()Found raster: cog
Opening: /vsis3/nasa-maap-data-store/file-staging/nasa-map/icesat2-boreal-v2.1/agb/0038382/boreal_agb_2020_202411261732637661_0038382.tif
CRS: PROJCS["unnamed",GEOGCS["GRS 1980(IUGG, 1980)",DATUM["unknown",SPHEROID["GRS80",6378137,298.257222101],TOWGS84[0,0,0,0,0,0,0]],PRIMEM["Greenwich",0],UNIT["degree",0.0174532925199433,AUTHORITY["EPSG","9122"]]],PROJECTION["Albers_Conic_Equal_Area"],PARAMETER["latitude_of_center",40],PARAMETER["longitude_of_center",180],PARAMETER["standard_parallel_1",50],PARAMETER["standard_parallel_2",70],PARAMETER["false_easting",0],PARAMETER["false_northing",0],UNIT["metre",1,AUTHORITY["EPSG","9001"]],AXIS["Easting",EAST],AXIS["Northing",NORTH]]
Bounds: BoundingBox(left=-1881478.0000000047, bottom=4653304.000000009, right=-1791478.0000000047, top=4743304.000000009)
Shape: 3000 x 3000
Bands: 2
Band 1 Stats:
Min: 0.43
Max: 11.79
Mean: 1.09
Band 2 Stats:
Min: 0.03
Max: 3.99
Mean: 0.20